Summary
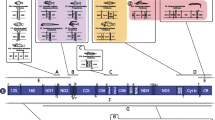
Sequence comparisons were made from 2214 bp of mitochondrial DNA cloned from six Pacific salmonid species. These sequences include the genes for ATPase subunit 6, cytochrome oxidase subunit 3, NADH dehydrogenase subunit 3, NADH dehydrogenase subunit 4L, tRNAGLY, and tRNAARG. Variation is found at 338 silent and 12 nonsilent positions of protein coding genes and 10 positions in the two tRNA sequences. A single 3-bp length difference was also detected. In all pairwise comparisons the sequence divergence observed in the fragment was higher than that previously predicted by restriction enzyme analysis of the entire molecule. The inferred evolutionary relationship of these species is consistent between methods. The distribution of silent variation shows a complex pattern with greatly reduced variation at the junctions of genes. The variation in the tRNA sequences is concentrated in the DHU loop. The close relationship of these species and extensive sequence analyzed allows for an analysis of the spectrum of substitutions that includes the frequencies of all 12 possible substitutions. The observed spectrum of substitutions is related to potential pathways of spontaneous substitution. The salmonid sequences show an extremely high ratio of silent to replacement substitutions. In addition the amino acid sequences of the four proteins coded in this fragment show a consistently high level of identity with theXenopus sequences. Taken together these data are consistent with a slower rate of amino acid substitution among the cold-blooded vertebrates when compared to mammals.
Similar content being viewed by others
References
Aquadro CF, Greenberg BD (1983) Human mitochondrial DNA variation and evolution: analysis of nucleotide sequences from seven individuals. Genetics 103:287–312
Atkinson T, Smith M (1985) Chapter 3. In: Gait MJ (ed) Oligonucleotide synthesis: a practical approach. IRL Press, New York, pp 35–81
Berg WJ, Ferris SD (1984) REstriction endonuclease analysis of salmonid mitochondrial DNA. Can J Fish Aquat Sci 41:1041–1047
Bibb MJ, Van Etten RA, Wright CT, Walberg MW, Clayton DA (1981) Sequence and gene organization of mouse mitochondrial DNA. Cell 26:167–180
Birnboim C, Doly J (1979) Rapid alkaline extraction procedure for screening recombinant plasmid DNA. Nucleic Acids Res 7:1513–1523
Brown AHD, Clegg MT (1983) Analysis of variation in related DNA sequences. In: Weir BS (ed) Statistical analysis of DNA sequence data. M Dekker, New York, pp 107–132
Brown GG, Simpson MV (1982) Novel features of animal mtDNA evolution as shown by sequences of two rat cytochrome oxidase subunit II gene. Proc Natl Acad Sci USA 79:3246–3250
Brown WM (1985) The mitochondrial genome of animals. In: MacIntyre RH (ed) Molecular evolutionary genetics. Plenum, New York, pp 95–130
Brown WM, George M Jr, Wilson AC (1979) Rapid evolution of animal mitochondrial DNA. Proc Natl Acad Sci USA 76:1967–1971
Brown WM, Prager EM, Wang A, Wilson AC (1982) Mitochondrial DNA sequences of primates: tempo and mode of evolution. J Mol Evol 18:225–239
Cann RL, Brown WM, Wilson AC (1984) Polymorphic sites and the mechanism of evolution in human mitochondrial DNA. Genetics 106:479–499
Clayton DA (1982) Replication of animal mitochondrial DNA. Cell 28:693–705
Clayton DA, Doda JN, Friedberg EC (1974) The absence of a pyrimidine dimer repair mechanism in mammalian mitochondria. Proc Natl Acad Sci USA 71:2777–2781
DeSalle R, Friedman T, Prager EM, Wilson AC (1987) Mitochondrial DNA sequences of HawaiianDrosophila. J Mol Evol 26:157–164
Fowler RG, Schaaper RM, Glickman BW (1986) Characterization of mutational specificity within thelac I gene for amutD5 mutator strain ofEscherichia coli defective in 3′ to 5′ exonuclease (proofreading) activity. J Bacteriol 167:130–137
Grantham R (1974) Amino acid difference formula to help explain protein evolution. Science 185:862–864
Gyllensten U, Wilson AC (1987) Mitochondrial DNA of salmonids: intraspecific variability detected with restriction enzymes. In: Ryman N, Utter FM (eds) The application of population genetics to fisheries management. University of Washington Press, Seattle, pp 301–317
Hattori M, Sakaki Y (1985) Dideoxy sequencing method using denatured plasmid templates. Anal Biochem 152:232–238
Henikoff S (1984) Unidirectional digestion with exonuclease III creates targeted breakpoints for DNA sequencing. Gene 28:351–359
Henikoff S (1987) Unidirectional digestion with exonuclease III in DNA sequence analysis. Meth Enzymol 155:156–165
Higuchi RG, Bowman B, Freiberger M, Ryder OA, Wilson AC (1984) DNA sequences from the quagga, an extinct member of the horse family. Nature 312:282–284
Higuchi RG, Wrischnik LA, Oakes E, George M, Tong B, Wilson AC (1987) Mitochondrial DNA of the extinct Quagga: relatedness and extent of postmortem change. J Mol Evol 25:283–287
Holmes DS, Quigley M (1981) A rapid boiling method for the preparation of bacterial plasmids. Anal Biochem 114:193–197
Lansman RA, Shade RO, Shapira JF, Avise JC (1981) The use of restriction endonucleases to measure mitochondrial DNA sequence relatedness in natural populations. J Mol Evol 17:214–226
Lindahl T, Nyberg B (1972) Rate of depurination of native deoxyribonucleic acid. Biochemistry 11:3610–3618
Loeb LA, Preston BD (1986) Mutagenesis by apurinic/apyrimidinic sites. Annu Rev Genet 20:201–230
Maniatis T, Fritsch EF, Sambrook J (1982) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratories, Cold Spring Harbor NY
Ojala D, Montoya J, Attardi G (1981) tRNA punctuation model of RNA processing in human mitochondria. Nature 290:470–474
Randall SK, Eritja R, Kaplan BE, Petruska J, Goodman MF (1987) Nucleotide insertion kinetics opposite abasic lesions in DNA. J Biol Chem 262:6864–6870
Roe BA, Ma D, Wilson RK, Wong JFH (1985) The complete nucleotide sequence of theXenopus laevis mitochondrial genome. J Biol Chem 260:9759–9774
Saccone S, Attimonelli M, Sbisa E (1985) Primary and higher order structural analysis of animal mitochondrial DNA. In: Quagliariello E, Slater C, Palmieri F, Saccone C, Kroon AM (eds) Achievements and perspectives of mitochondrial research. Elsevier/North-Holland, Amsterdam, pp 37–47
Schaaper RM, Dunn RL (1987) Spectra of spontaneous mutations inEscherichia coli strains defective in mismatch correction: the nature ofin vivo DNA replication errors. Proc Natl Acad Sci USA 84:6220–6224
Takeshita M, Chang C, Johnson F, Will S, Grollman AP (1987) Oligodeoxynucleotides containing synthetic abasic sites: model substrates for DNA polymerases an apurinic/apyrimidinic endonucleases. J Biol Chem 262:10171–10179
Thomas WK, Withler RE, Beckenbach AT (1986) Mitochondrial DNA analysis of Pacific salmonid evolution. Can J Zool 64:1058–1064
Tomkinson AE, Linn S (1986) Purification and properties of a strand-specific endonuclease from mouse cell mitochondria. Nucleic Acids Res 14:9579–9593
Utter FM, Allendorf FW, Hodgins HO (1973) Genetic variability and relationships in Pacific salmon and related trout based on protein variations. Syst Zool 22:258–270
Wilson AC, Cann RL, Carr SM, George M, Gyllensten UB, Helm-Bychowski KM, Higuchi RG, Palumbi SR, Prager EM, Sage RD, Stoneking M (1985) Mitochondrial DNA and two perspectives on evolutionary genetics. Biol J Linn Soc 26:375–400
Wolstenholme DR, Macfarlane JL, Okimoto R, Clary DO, Wahleithner JA (1987) Bizarre tRNAs inferred from DNA sequences of mitochondrial genomes of nematode worms. Proc Natl Acad Sci USA 84:1324–1328
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Thomas, W.K., Beckenbach, A.T. Variation in salmonid mitochondrial DNA: Evolutioinary constraints and mechanisms of substitution. J Mol Evol 29, 233–245 (1989). https://doi.org/10.1007/BF02100207
Received:
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF02100207