- Split View
-
Views
-
Cite
Cite
Christian Schmitz-Linneweber, Ralph Regel, Tung Gia Du, Holger Hupfer, Reinhold G. Herrmann, Rainer M. Maier, The Plastid Chromosome of Atropa belladonna and its Comparison with that of Nicotiana tabacum: The Role of RNA Editing in Generating Divergence in the Process of Plant Speciation, Molecular Biology and Evolution, Volume 19, Issue 9, September 2002, Pages 1602–1612, https://doi.org/10.1093/oxfordjournals.molbev.a004222
Close - Share Icon Share
Abstract
The nuclear and plastid genomes of the plant cell form a coevolving unit which in interspecific combinations can lead to genetic incompatibility of compartments even between closely related taxa. This phenomenon has been observed for instance in Atropa-Nicotiana cybrids. We have sequenced the plastid chromosome of Atropa belladonna (deadly nightshade), a circular DNA molecule of 156,688 bp, and compared it with the corresponding published sequence of its relative Nicotiana tabacum (tobacco) to understand how divergence at the level of this genome can contribute to nuclear-plastid incompatibilities and to speciation. It appears that (1) regulatory elements, i.e., promoters as well as translational and replicational signal elements, are well conserved between the two species; (2) genes—including introns—are even more highly conserved, with differences residing predominantly in regions of low functional importance; and (3) RNA editotypes differ between the two species, which makes this process an intriguing candidate for causing rapid reproductive isolation of populations.
Introduction
The compartmented genetic system of the plant cell is of heterogenous phylogenetic origin. Plastids and mitochondria are acquisitions of cyanobacterial and α-proteobacterial ancestry (Gray 1999 ), whereas the origin of the nucleus, although showing a number of archebacterial characteristics, is still in debate (Ribeiro and Golding 1998 ; Bell 2001 ). It has been speculated that coadaptation of these cellular genetic compartments contributes to speciation processes (Caspari 1948 ; Michaelis 1954 ; Stubbe 1989 ) by impairing nucleus-plastid interaction in interspecific crosses as has been described in detail in the genus Oenothera (Stubbe 1989 ; Chiu and Sears 1993 ). The progeny of these crosses can exhibit severe developmental disturbances, designated hybrid bleaching, or hybrid variegation (Renner 1929 , 1934 ). This kind of incompatibility has been observed with various materials (Kirk and Tilney-Bassett 1987 ), including somatically generated hybrids, the so-called cybrids, which often display developmental defects like chlorophyll deficiency, disordered thylakoid membranes, and dwarfism (Pelletier et al. 1983 ; Pental et al. 1986 ; Kushnir et al. 1991 ). It has been shown that the severity of the defects increases with phylogenetic distance of the species involved. For instance, cybrids between Nicotiana tabacum and Nicotiana plumbaginifolia seem to be perfectly healthy (Menczel et al. 1986 ), whereas transfer of plastids between distantly related Solanacean species of one genus (Perl, Aviv, and Galun 1991 ) or different Solanacean genera (Than et al. 1988 ) can have deleterious consequences. A particular interesting case is the transfer of plastids between deadly nightshade (Atropa belladonna) and tobacco (N. tabacum). Both species are also members of the Solanaceae but belong to different tribes, the Solaneae and Cestreae, respectively. Here, the Atropa plastid in the tobacco nuclear background develops normally, whereas the reverse combination is white and grows only heterotrophically (Kushnir et al. 1987 , 1991 ). The obvious reduction in fitness of this artificial cybrid as well as many others and hybrids suggests that compromised nucleus-plastid interactions contribute to reproductive barriers in natural populations, too. An unsettled task is the determination of loci shaping these barriers.
The coding potential of plastids, the plastome, is deposited in a multitude of identical circular DNA molecules of 75–250 kbp, depending on the plant species. Sequences of plastid chromosomes are available for a variety of vascular plants and algae (for a list of completed plastid chromosomes, see http://www.ncbi.nlm.nih.gov/PMGifs/Genomes/plastids_tax.html). Their analysis has so far focused on the question of coding potential and the origin of plastids and land plants (Martin et al. 1998 ; Gray 1999 ; Stoebe and Kowallik 1999 ; Lemieux, Otis, and Turmel 2000 ). Available sequence data have revealed the conservative nature of contemporary plastid chromosomes with regard to both structure and gene content. The similarity is particularly striking when defined phylogenetic groups are considered, for example, the streptophytes and their sister group, the chlorophytes (http://www.ncbi.nlm.nih.gov/PMGifs/Genomes/plastids_tax.html). With rare exceptions, these plastid chromosomes exhibit a quadripartite structure and contain a similar set of genes, specifying predominantly constituents of the photosynthetic and genetic machineries. Genes and operons are densely packed, with conserved reading frames sometimes even overlapping; intergenic regions are relatively short and often target sites for nuclear-coded factors regulating plastid gene expression and implicit development. For instance, nuclear-coded sigma-factors mediate transcription initiation by the plastid-coded core RNA polymerase (Allison 2000 ), and nuclear-coded proteins bind to 5′ or 3′ untranslated regions (UTRs) of plastid messages that influence their turnover rates (Monde, Schuster, and Stern 2000 ). Furthermore, plastid messages, most of them polycistronic, have to be dissected, trimmed, polyadenylated, spliced, and edited, all of which depends on specific interactions with nuclear factors that have to be imported from the cytoplasm (Barkan and Goldschmidt-Clermont 2000) .
Sequence comparisons of complete plastid chromosomes have been carried out using exclusively more distantly related lineages, except the Graminean species rice, maize, and wheat (Maier et al. 1995 ; Ogihara et al. 2002 ), for which hot-spots of divergence have been analyzed with respect to mechanistic rather than evolutionary issues. Aspects of microevolution and their impact on plant speciation have been addressed only by comparisons of a few plastid sequence stretches (e.g., Young and dePamphilis 2000) . From these data, it is clear that differences reside predominantly in the rapidly evolving intergenic regions when closely related plants are analyzed (e.g., Gielly and Taberlet 1994 ). Intergenic regions harbor many sequence elements functioning in gene expression like promoters or translational enhancers, which are indispensable for development and maintenance of the organelle. In general, these elements are far more conserved than the surrounding sequence (e.g., Manen, Savolainen, and Simon 1994 ). The question remains whether it is the divergence of intergenic functional elements and hence the regulation of gene expression that causes incompatibility of genetic compartments, or whether the coding regions themselves are the source of this phenomenon.
Here, we present the complete sequence of the plastid chromosome of A. belladonna and compare it with the corresponding sequence of tobacco (Shinozaki et al. 1986 ), with emphasis on differences relevant to interspecific incompatibilities of plastid and nuclear genomes.
Materials and Methods
Leaf material from A. belladonna Ab5, a transgenic kanamycin-resistant line (Kushnir et al. 1991 ), was harvested 4 weeks after germination. Chloroplasts were isolated by Percoll gradient centrifugation as described previously (Robinson and Barnett 1988 ). DNA was isolated using a CTAB-based protocol (Murray and Thompson 1980 ) and RNA with TRIzol Reagent (GIBCOBRL, Eggenheim).
RNA was reverse transcribed with a random primer mixture using Superscript™ (GIBCOBRL; Maier et al. 1995 ). DNA and cDNA were amplified according to a standard protocol with 30 cycles of 94°C (for 20 s), 55°C (for 20 s), and 72°C (for 45 s) with a 2-min extension at 94°C for the first cycle and a 4-min extension at 72°C for the final cycle in the presence of 2.5 mM magnesium chloride.
A recombinant DNA library containing XhoI, BamHI, SalI, PstI, and ApaI plastid DNA fragments of A. belladonna Ab5 in pBluescript II SK- (Stratagene, La Jolla, CA) provided templates for sequencing. PCR-derived amplification products were used to close gaps between restriction fragments. Nucleotide sequences were determined by the dideoxy chain termination method (Sanger, Nicklen, and Coulsen 1977 ) using a set of approximately 400 representative primers for both DNA strands with an ABI 377 sequencer (Applied Biosystems, USA). Oligonucleotides used for the PCR and for sequencing were purchased from MWG Biotech (Ebersberg, Germany). Sequences were evaluated and assembled using Sequencher 3.0 (Gene Codes Corporation, USA). The sequences were aligned using the BioEdit sequence alignment editor (North Carolina State University).
Results and Discussion
Structure and Gene Content of the Atropa Plastid Chromosome in Comparison to that of Tobacco
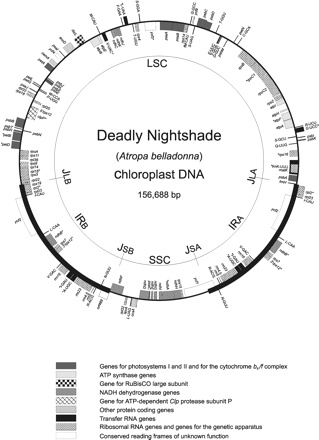
The Atropa ptDNA sequence assembles as a circle of 156,688 bp (accession number: AJ316582; tobacco: 155,939 bp), with an overall G + C content of 37.6%. A large inverted repeat (IR; 25,906 bp) containing the rRNA operon separates a large single copy (LSC; 86,868 bp) and a small single copy (SSC; 18,008 bp) region (fig. 1 ). The overall similarity of the two DNA molecules is greater than that of any previously sequenced pair of ptDNAs, exceeding 96% identity. The Atropa plastid chromosome, like that of tobacco, encodes a total of 113 identified genes arranged in an identical order along the plastid chromosome in the two species. This includes the presence of sprA, a gene encoding an RNA suggested to influence 16S rRNA maturation (Vera and Sugiura 1994 ; Sugita et al. 1997 ), and so far found only in tobacco and tomato, which is 94% identical in Atropa. On the other hand, infA, a gene coding for a translation initiation factor in other plant species, is a pseudogene in both tobacco and Atropa. Another known peculiarity of the tobacco plastid chromosome, distinguishing it from other dicot plants, concerns the overlap of the rpl22 and rps3 reading frames. This overlap, as well as all other gene overlaps found in tobacco, is also seen in Atropa. Intron numbers and positions are highly conserved. The two species share the same set of introns: altogether 20 group II introns and 1 group I intron.
In tobacco, in addition to known genes, 11 open reading frames (ORFs) of unknown function, which are at least 70 codons in length, have been annotated. Six of them are conserved in Atropa, probably because all but one of them (ORF99) map to the IR (table 1 ), which is known to evolve far less rapidly than the single copy regions (Maier at al. 1995 ; Sugiura 1995 ). The five remaining tobacco ORFs have no adequate counterpart in Atropa, terminating earlier either because of deletions or point mutations. The finding that these ORFs are not conserved in Atropa corroborates previous data questioning their significance (Schmitz-Linneweber et al. 2001a ).
The Atropa plastid chromosome is 749 bp longer than its tobacco counterpart. This size difference is largely on account of an extension of the IR region that reaches 542 bp farther into the SSC region than the tobacco IR. Contraction and expansion of IRs in chloroplast chromosomes are well known from various other species (Maier et al. 1990 ; Goulding et al. 1996 ). In addition, many insertions and deletions (indels) contribute to the size difference between the Atropa and tobacco chromosomes. Relative to tobacco, there are 65 insertions and 60 deletions equal to or larger than 5 bp in intergenic regions and in introns of the plastid chromosome of Atropa; approximately half of them are associated with direct repeats (data not shown). The latter repeats may reflect replication slippage or misaligning events during recombination (or both) (Hancock 1996 ). Most of the observed repeats reside in AT-rich regions. In bacteria, such regions have been shown to be particularly prone to replication slippage (Levinson and Gutman 1987 ). It has been shown earlier that direct repeats contribute significantly to variability of plastid chromosomes even between closely related species (Aldrich et al. 1988 ; Wolfe, Morden, and Palmer 1992 ).
Coding Regions are Highly Conserved Between Atropa and Tobacco
As an important interface in nuclear-plastid interactions, plastome-encoded polypeptides and RNAs, as well as distinct sequence motifs in the plastid chromosome itself have to interact with nuclear-encoded polypeptides (and possibly RNAs), for instance during gene expression or assembly of multiprotein complexes. Considering that coevolution of polypeptides with their interaction partners is a well-described phenomenon (Goh et al. 2000) , divergence of plastid polypeptides may hamper interspecific interactions of compartments. Differences in homologous polypeptide sequences encoded by the Atropa and tobacco plastid chromosomes occur as amino acid substitutions and, rarely, also as insertions and deletions.
Altogether 10 indels are present in six polypeptide genes, of which six can be attributed to replication slippage (fig. 2 and data not shown). The eight codons not present in AtropaaccD, which encodes the β-subunit of the prokaryotic-type acetyl-CoA–carboxylase, reside in a region of relatively low sequence conservation among land plant species (fig. 2A ). Deletions relative to tobacco occur in most other species examined. Similarly, a single codon missing in AtropaccsA, specifying a protein involved in c-type cytochrome synthesis, is located in a short stretch coding for rather poorly conserved amino acid sequences (fig. 2B ). In AtroparpoC2, coding for a subunit of a plastid RNA polymerase, three successive codons are absent relative to tobacco and all other higher plant sequences examined. Deletions further downstream affecting the same region of RpoC2 are also known from other plant species (fig. 2C ). For Atroparps16, specifying a ribosomal polypeptide, a 9-bp deletion including the tobacco stop codon leads to a short carboxy-terminal extension of the encoded polypeptide identical to the one found in spinach (Spinacia oleracea). It has been shown that the Rps16 C-terminus does not interact with rRNA or with other ribosomal proteins, which questions its importance in ribosomal function (Allard et al. 2000 ) (fig. 2D ). Additional indels have been found in the ycf1 and ycf2 reading frames. These two genes are known to be essential for plant survival (Drescher et al. 2000) , though their functions have not been determined yet. Unlike ycf2,ycf1 is speckled with indels and rarely occurring conserved elements. None of the four deletions noted in Atropaycf1 affects these conserved elements, and all of the codons deleted in tobacco ycf1 are also absent in other dicot species (data not shown). For ycf2, the two insertions found in tobacco, one of them three codons in size, the other eight, form direct repeats with adjacent sequence elements and are also situated outside highly conserved domains (data not shown).
In Atropa, rrn16 has two insertions of four and five nucleotides relative to tobacco, spaced by three base pairs. The region is conserved in vascular plants, however, in algae, it is a hot spot of divergence (fig. 2E ). Neither rRNA-ribosomal protein interactions nor any other interactions (with initiation factors, tRNAs, mRNA, or antibiotics) have been noted for the homologous region in Escherichia coli (Mueller and Brimacombe 1997a , 1997b ), which questions the importance of this domain. In summary, all indels in the coding regions of the Atropa and tobacco plastid chromosomes are located outside conserved domains and therefore seem to have no decisive impact on protein structure and function.
Most amino acid substitutions caused by differences in the plastome sequences of Atropa and tobacco occur at nonconserved positions in polypeptides or are conservative, that is, probably without effect on protein function. But 11 of the altogether 360 substitutions (∼3%) concern highly conserved residues, a phenomenon affecting the products of nine genes (table 2 ). To assess the potential functional importance of these residues, we analyzed conservation of the sequence neighborhood by counting variable residues in a window of 20 amino acids surrounding the residue in question. The position of the window was chosen to minimize the number of variable positions, as the residue of interest can be at the edge as well as in the center of a conserved domain. The proportion of all variable positions in this window yields a value suitable for comparison of the impact on domain conservation of different substitutions (table 2 ). Among the five substitutions located in the domains most rigidly conserved, four are reverted by C-to-U editing of the respective codon in the tobacco transcript (at atpA site 1, the psbE site and rps14 site 1, according to Tsudzuki, Wakasugi, and Sugiura [2001] , and at one site in ndhD, here referred to as ndhD site 3, not described so far [unpublished data]). The fifth, found in PsaB, a core subunit of photosystem I, is located at a surface region of the protein that is suggested not to interact with neighboring components (Jordan et al. 2001) , as revealed by analysis of structural data of the homologous polypeptide in Synechococcus (PDB Id: 1JB0) using SWISS PDB viewer V3.7b2 (www.expasy.ch/spdbv/mainpage.html). Similarly, for two substitutions at less-conserved positions in AtpB, the β-subunit of ATP synthase, residues are found exposed to solvent (PDB IFX0; Groth and Pohl 2001) . In contrast, the L264 in Atropa AtpA, a component of the same multiprotein complex, is situated at the core of the protein and in silico exchange to proline, the unedited state in tobacco, revealed undesirable interactions with A267 and F318 (data not shown). In summary, most of the codon substitutions at highly conserved positions are rescued by RNA editing, so that differences on the chromosomal level are adjusted on the transcript level. The reverse, that diverse amino acid positions result from differences in editing, has not been observed.
Known Regulatory Modules from Tobacco are Almost Identical in Atropa
Promoters
Two principal promoter classes have been identified that operate with two distinct RNA polymerases, one nuclear (NEP), the other plastid encoded (PEP; Weihe and Börner 1999 ). The latter enzyme recognizes prokaryotic-type promoters that are usually furnished with “−10 and −35” boxes. Of the 29 transcription initiation sites identified in tobacco, 16 have been assigned to this class according to their preferential usage in wild-type but not in PEP-deficient material, whereas the remaining 13 promoters fall into the NEP class (Liere and Maliga 1999 and references therein; Allison 2000 ).
All −10 and −35 boxes of PEP promoters as well as all YAT-motives in NEP promoters are identical between Atropa and tobacco. In addition, elements regulating promoter activity, like those upstream of the psbD blue-light promoter or the rpoB promoter (Allison and Maliga 1995 ; Inada et al. 1997 ), are identical. Base substitutions between Atropa and tobacco promoters have only been found in four instances (trnG-UCC, atpI,rps16, and clpP), but always outside the core elements. In E. coli, it has been shown that sequences outside the −10 and −35 boxes may be randomized without compromising promoter function (Busby and Ebright 1994 ), and for NEP promoters, mutational analysis have shown that mutations outside the YAT motif decrease, but do not abolish transcriptional activity (Kapoor and Sugiura 1999 ; Liere and Maliga 1999 ). Collectively, promoter structures are almost invariant between the two plastid chromosomes. Concerning nuclear trans factors necessary for transcription of plastid genes, it has been shown that they can differ between higher order taxa, leading to a differential usage of one of several distinct promoters situated upstream of a single transcription unit (Sriraman, Silhavy, and Maliga, 1998 ). According to these data, the apparatus of both species should be capable of properly initiating transcription in a given promoter region in organelle exchange situation, although initiation sites may differ.
Shine-Dalgarno Sequences and Other Translation-Mediating Elements
Sequence requirements for plastid translation are relatively poorly characterized (Zerges 2000) . The plastid chromosome codes for only a limited number of ribosomal proteins and depends on nuclear factors for both mechanistic and regulatory aspects of translation. Only few of these regulatory nuclear factors are known from higher plants (Sugiura, Hirose, and Sugita 1998 ; Fisk, Walker, and Barkan 1999 ). Although the plastid translational machinery is of prokaryotic origin, quite a few plastid genes do not have any obvious Shine-Dalgarno (SD)-like sequence, and if they do, it is often shifted to a more upstream position, untypical for eubacterial translation systems (Sugiura, Hirose, and Sugita 1998 ). In some instances, novel sequence elements not found in prokaryotes seem to replace or extend the function of SD sequences (e.g., Hirose and Sugiura 1996 ). Comparison of all known sequence elements for tobacco plastid translation with the homologous sequences in Atropa revealed either complete (all SD-like sequences) or nearly complete (other translational enhancers) identity (data not shown). It therefore appears unlikely that they have played a significant role in generating genetic diversity of these two species.
Origins of Replication
Origins of plastome replication are known to be divergent between species examined so far (Kunnimalaiyaan and Nielsen 1997 ). It has been proposed that differences in origins of replication between different species affect the specificity and efficiency of replication (Hornung et al. 1996 ). In tobacco, two replication origins have been mapped using electron microscopy of replication intermediates and an in vitro replication approach (Kunnimalaiyaan and Nielsen 1997 ; Kunnimalaiyaan, Shi, and Nielsen 1997 ). One of them, designated OriA, is situated within the trnI-GAU intron. It consists minimally of 82 bp with a stem-loop structure and a direct repeat. Only one base is changed in Atropa relative to tobacco. The second element, termed OriB, was mapped to the ycf1 gene. It also comprises a large stem-loop structure and several direct repeats. In this case, 4 out of 243 bp differ between Atropa and tobacco. A third sequence element conferring autonomous replication to plasmids in yeast has been described in tobacco situated within the ndhF reading frame (Ohtani et al. 1984 ). This element comprises approximately 350 bp, of which only five differ between Atropa and tobacco. Differences in replication efficiency have been invoked in Oenothera to play a role in the segregation rates of different plastid types combined in one cell (Hornung et al. 1996 ). It is known that most, if not all, components of the replication machinery are coded in the nucleus. In the genus Oenothera, species differ substantially in the number of direct repeats and overall length of the region containing the replication origin (Sears, Stoike, and Chiu 1996 ). Interestingly, tobacco and Atropa exhibit no substantial divergence in their predicted Ori structures. It therefore seems unlikely that differences in replication arising from alterations in the three origins identified so far in tobacco are responsible for the observed compartmental incompatibility in cybrids between Atropa and tobacco.
Introns and Editing Sites
Before translation, plastid messages can be subject to RNA splicing and RNA editing, two posttranscriptional processing events that vitally interfere with the message's coding content (Barkan and Goldschmidt-Clermont 2000 ; Bock 2000 ). Intron structures are highly conserved between Atropa and tobacco. The crucial determinants for intron splicing, that is, the splice donor and acceptor sites, are identical, with five exceptions. The clpP second intron–third exon boundary exhibits a 5-bp deletion in Atropa (fig. 3 ). Furthermore, single base pair substitutions occur between Atropa and tobacco in the intron–3′exon boundary region of trnL(UAA), trnV(UAC), rps16 and the second intron of ycf3 (fig. 3 ). These differences impartially affect introns of different classes, suggesting that there is no biased evolution of a subgroup of introns in the two species. It remains a matter of speculation whether these differences might affect splicing of the respective genes in interspecific combinations of organelles, but it is for certain noteworthy that all four mentioned differences in group II introns affect domain VI. Domain VI is the most rigidly conserved part of the whole intron structure (Michel, Umesono, and Ozeki 1998 ) and is central to the splice chemistry. Interactions of nuclear factors with domain VI may therefore contribute to compartmental incompatibilities.
RNA editing leads to changes of single C to U residues, sometimes restoring initiation codons, but more often internal codons of high functional relevance (Bock 2000 ). The editotype of tobacco, that is, all editing sites found in the complete transcriptome, has been determined previously (Hirose et al. 1999 ) and described to consist of 31 sites scattered throughout the plastid chromosome. Of these sites, 28 are conserved in Atropa, whereas sites in rps14 (site 2), psbE, atpA (site 1) (designation of sites according to Tsudzuki, Wakasugi, and Sugiura 2001) , and a newly identified site in tobacco ndhD, here referred to as site 3 (position 118352 of the tobacco plastid chromosome, accession no. Z00044; unpublished data), are absent, i.e., a T-residue is already found at the respective positions in Atropa DNA.
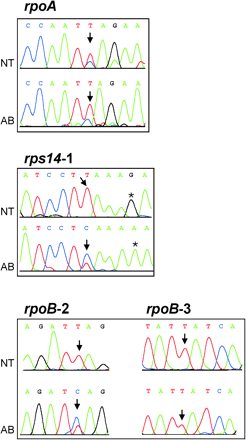
To determine whether Atropa is capable of processing the 28 shared sites, corresponding cDNA fragments were amplified by polymerase chain reaction (PCR) and sequenced directly. It turned out that Atropa harbors activities to edit all 28 sites, although with two quantitative differences to tobacco: both rpoB (site 2) and rps14 (site 1) are edited to a lesser extent in Atropa tissue in comparison with tobacco (fig. 4 ). In both cases, editing efficiency, which is 100% in tobacco, is greatly reduced in Atropa, with the signal from the unedited message, the C-peak, even predominating. Other editing sites are fully edited in Atropa, including, for instance, rpoB site 3 from the same message (fig. 4 ), indicating that contaminating DNA is not the cause of the observed C-signal. In addition, sites known to be only partially edited, like ndhD site 1 and rpoA, are processed to a similar extent in both species (fig. 4 and data not shown).
Atropa-Specific Editing Sites
To determine sites specific to Atropa but absent from tobacco, interplastomic codon changes that could potentially be reverted by RNA editing of Atropa messages were checked by cDNA sequencing. This screen was carried out on the basis of all known codon transitions occurring in plastids of angiosperms as a consequence of RNA editing (Tsudzuki, Wakasugi, and Sugiura 2001) . In this way, three sites present in Atropa but absent from tobacco were found in the genes ndhA (codon 189), ndhD (codon 293), and rpoB (codon 809) (not shown). All three editing events mediate serine to leucine codon changes. Homologs of the ndhA and ndhD site are known from monocots, whereas the rpoB site has not been described in other species.
What evolutionary or functional significance might be attributed to these findings? First, our data emphasize that RNA editing is a rapidly evolving trait, with editing sites changing quickly between taxa (Freyer, Kiefer-Meyer, and Kössel 1997 ; Corneille, Lutz, and Maliga 2000 ; Tsudzuki, Wakasugi, and Sugiura 2001 ). Second, the reduced editing efficiency found for rps14 (site 1) and rpoB (site 2) might be interpreted as the first step on the way to losing an editing site because a marginally edited site ultimately poses the danger of incorporating the “wrong” amino acid into the respective protein. This should launch a counterselective pressure favoring C-to-T transitions already on DNA, thereby obliterating the need to edit the site at all and hence relieving the problem caused by partial editing. In this scenario, a decreased editing efficiency would be a dysfunctional chance event entailing the loss of a site, but it is of course also conceivable that both messages, the edited as well as the unedited, serve some purpose, maybe because both amino acids are beneficial in different ways for protein function (Hirose et al. 1999 ; Karcher and Bock 2002 ).
Conclusions
Understanding the molecular basis for the divergence of species is a central aspect of current biology (Barton 2001) . In comparing the plastid chromosomes of two closely related species from which nuclear-plastid cybrids with impaired development can be somatically constructed, we attempted to identify genetic determinants of plastids relevant for speciation. Of course, microdiversity resulting in speciation can be a gradual process, and therefore relies on all kinds of differences in an additive way. This is corroborated by studies on hybrid incompatibilities, for which multigenic, epistatic processes seem to play a major role in speciation (Orr 1994 ). On the other hand, the influence of “major genes” in speciation is also discussed (Orr 2001 ).
It had previously been noted that intergenic sequence is a source for interspecific variance (Wolfe, Li, and Sharp 1987 ; Gielly and Taberlet 1994 ). It was therefore tempting to speculate that the regulatory elements responsible for plastid gene expression, which reside predominantly in intergenic regions, are rapidly evolving and play a key role in interspecific divergence. However, in the intrafamilial comparison of the two plastid chromosomes presented here, promoters, translational enhancers, and replication origins are well conserved, as are the coding regions. In contrast, RNA editotypes differ between the two species both qualitatively and quantitatively. RNA editing thus fulfils all requirements for a major trait in speciation because it is evolving rapidly and may have a substantial effect on plant fitness via protein function (Bock, Kössel, and Maliga 1994 ; Zito et al. 1997 ; Sasaki et al. 2001 ). Differences in editotypes in this light have the quality of a yes or no decision with regard to protein function, quite in contrast to the accumulative effect of other differences described here. Because the nuclear-encoded editing machinery is known to be species- and site-specific (Chaudhuri, Carrer, and Maliga 1995 ; Bock and Koop 1997 ; Schmitz-Linneweber et al. 2001b ), it is imperative for a plant to express the correct set of editing factors. It has to be expected that whenever an editotype from one species is joined with an inadequate set of nuclear factors of a second one, as in interspecific hybrids and cybrids, editing failures and thus fatal defects will ensue. However, this does not explain the marked differences in Atropa-tobacco and tobacco-Atropa cybrid phenotypes. As our data show, the nuclei of both cybrids, the tobacco as well as the Atropa, face the challenge of editing sites not present in their genuine plastome. But only recently we could show that the allotetraploid nucleus of tobacco harbors more editing factors than needed for the endogenous sites and is therefore capable of heterologous editing. Interestingly, this has been shown for a spinach site, which is homologous to the Atropa-specific ndhA site described here and which is edited efficiently in the tobacco nuclear background (Schmitz-Linneweber et al. 2001b ). Hence, it is expected that the tobacco nucleus is also capable of editing at least this Atropa-specific site.
Conclusively, we propose that plastid chromosomal divergence affects speciation in two aspects. First, a broad variety of differences in polypeptides or intron structure generates gradual deviation between populations. Second, RNA editing as a trait with decisive impact on plant fitness might influence speciation in a more saltatorial manner. Work on interspecific hybrids and cybrids should help us to understand which of these components of divergence is most influential in producing new species.
Geoffrey McFadden, Reviewing Editor
Keywords: sequence analysis plastid chromosome Atropa belladonna nucleus-plastid incompatibility speciation
Address for correspondence and reprints: Botanisches Institut der Ludwig-Maximilians-Universität München, Menzinger Str. 67, 80638 München, Germany. E-mail: raimaier@botanik.biologie.uni-muenchen.de
Table 1 Comparison of the 11 Annotated ORFs from Tobacco with Homologous Regions in Atropa
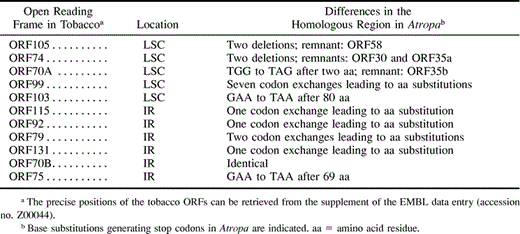
Table 1 Comparison of the 11 Annotated ORFs from Tobacco with Homologous Regions in Atropa

Table 2 Substitutions of Conserved Amino Acids Between Tobacco and Atropa in All Known Protein Coding Plastid Genes due to Point Mutations in the Corresponding Codons
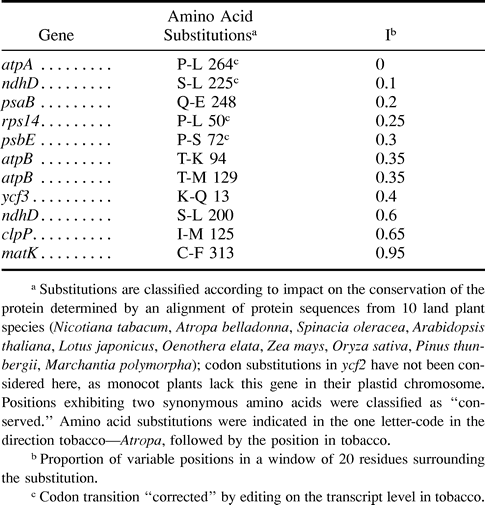
Table 2 Substitutions of Conserved Amino Acids Between Tobacco and Atropa in All Known Protein Coding Plastid Genes due to Point Mutations in the Corresponding Codons

Fig. 1.—Gene organization of the circular A. belladonna plastid chromosome. Large and small single copy regions (LSC, SSC) are separated by the inverted repeats IRA and IRB (bold), respectively. Intron-containing genes are marked by asterisks. Genes drawn inside the circle are transcribed clockwise, those outside anticlockwise. ORF489 located at the IRB-SSC border represents a truncated form of ycf1. Genes belonging to different functional groups are marked by different gray-scales or patterns (see legend below). For abbreviations and nomenclature of protein-coding genes, see Stoebe, Martin, and Kowallik (1998)
Fig. 2.—Alignment of four plastid amino acid sequences and one plastid nucleotide sequence exhibiting indels between tobacco and Atropa. Sequence elements deleted between Atropa and tobacco are marked in boldface letters. Identities to the tobacco sequence are indicated by dots. Numbers refer to positions in the tobacco protein and rRNA gene, respectively. NT, Nicotiana tabacum; AB, Atropa belladonna; AT, Arabidopsis thaliana, OE, Oenothera elata; LJ, Lotus japonicus; SO, Spinacia oleracea; EV, Epifagus virginiana; PT, Pinus thunbergii; MP, Marchantia polymorpha; ZM, Zea mays; OS, Oryza sativa, GT, Guillardia theta, OSI, Odontella sinensis, PP, Porphyra purpurea; CP, Cyanophora paradoxa; EG, Euglena gracilis; CV, Chlorella vulgaris. Sequences were obtained via http://www.ncbi.nlm.nih.gov/PMGifs/Genomes/plastids_tax.html.
Fig. 3.—Alignment of intron-exon boundaries with differences between tobacco and Atropa. Sequence stretches spanning intron-exon boundaries were aligned as described previously (Sugita and Sugiura 1996 ). Sequence differences are marked in bold. Introns are classified according to a previously established system (Michel, Umesono, and Ozeki 1998 ). Upper sequence, N. tabacum (EMBL accession no. Z00044). Lower sequence, A. belladonna (EMBL accession no. AJ316582)
Fig. 4.—Quantitative editing differences between Atropa and tobacco. Specific cDNAs of Atropa and tobacco were amplified and sequenced directly. Relevant short intervals of a few nucleotides are shown for three cDNAs (rpoA, rps14, rpoB) together with the corresponding chromatograms. The editing sites are marked by arrows. The two rpoB editing sites shown originate from the same sequencing reaction (numbers refer to previously established nomenclature (Hirose et al. 1999 ). A base substitution 3′ to the editing site in rps14 is denoted by asterisks
We thank Ingrid Duschanek for skillful technical assistance and an anonymous reviewer for helpful comments on the manuscript. This work was supported by the Deutsche Forschungsgemeinschaft (SFB 184 and TR1) and the Fonds der Chemischen Industrie.
References
Aldrich J., B. W. Cherney, E. Merlin, L. Christopherson,
Allard P., A. V. Rak, B. T. Wimberly, W. M. Clemons Jr,, A. Kalinin, M. Helgstrand, M. B. Garber, V. Ramakrishnan, T. Hard,
Allison L. A., P. Maliga,
Barkan A., M. Goldschmidt-Clermont,
Bell P. J. L.,
Bock R.,
Bock R., H. U. Koop,
Bock R., H. Kössel, P. Maliga,
Busby S., R. H. Ebright,
Chaudhuri S., H. Carrer, P. Maliga,
Chiu W. L., B. B. Sears,
Corneille S., K. Lutz, P. Maliga,
Drescher A., S. Ruf, T. Jr Calsa, H. Carrer, R. Bock,
Du Y. C., S. Hong, R. J. Spreitzer,
Fisk D. G., M. Walker, A. Barkan,
Freyer R., M.-C. Kiefer-Meyer, H. Kössel,
Gielly L., P. Taberlet,
Goh C. S., A. A. Bogan, M. Joachimiak, D. Walther, F. E. Cohen,
Goulding S. E., R. G. Olmstead, C. W. Morden, K. H. Wolfe,
Groth G., E. Pohl,
Hirose T., T. Kusumegi, T. Tsudzuki, M. Sugiura,
Hirose T., M. Sugiura,
Hornung S., H. Fulgosi, P. Dorfel, R. G. Herrmann,
Inada H., M. Seki, H. Morikawa, M. Nishimura, K. Iba,
Jordan P., P. Fromme, H. T. Witt, O. Klukas, W. Saenger, N. Krauß,
Kapoor S., M. Sugiura,
Karcher D., R. Bock,
Kirk J. T. O., R. A. E. Tilney-Bassett,
Koop H. U., K. Steinmüller, H. Wagner, C. Rößler, C. Eibl, L. Sacher,
Kunnimalaiyaan M., B. L. Nielsen,
Kunnimalaiyaan M., F. Shi, B. L. Nielsen,
Kushnir S. G., E. Babiychuk, M. Bannikova, V. Momot, I. Komarnitsky, N. Cherep, Y. Gleba,
Kushnir S. G., L. R. Shlumukov, N. J. Progebnyak, S. Berger, Y. Gleba,
Lemieux C., C. Otis, M. Turmel,
Levinson G., G. A. Gutman,
Liere K., P. Maliga,
Maier R. M., I. Dory, G. Igloi, H. Kössel,
Maier R. M., K. Neckermann, G. L. Igloi, H. Kössel,
Manen J. F., V. Savolainen, P. Simon,
Martin W., B. Stoebe, V. Goremykin, S. Hansmann, M. Hasegawa, K. Kowallik,
Menczel L., L. S. Polsby, K. E. Steinback, P. Maliga,
Michaelis P.,
Michel F., K. Umesono, H. Ozeki,
Monde R. A., G. Schuster, D. B. Stern,
Mueller F., R. Brimacombe,
———.
Murray M. G., W. F. Thompson,
Ogihara Y., K. Isono, T. Kojima, et al. (16 co-authors)
Ohtani T., H. Uchimiya, A. Kato, H. Harada, M. Sugita, M. Sugiura,
Orr H. A.,
Pelletier G., C. Primard, F. Vedel, P. Chetrit, R. Remy, Rousselle, M. Renard,
Pental D., J. D. Hamill, A. Pirrie, E. C. Cocking,
Perl A., D. Aviv, E. Galun,
Renner O.,
———.
Ribeiro S., G. B. Golding,
Robinson C., L. K. Barnett,
Sanger F., S. Nicklen, A. R. Coulsen,
Sasaki Y., A. Kozaki, A. Ohmori, H. Iguchi, Y. Nagano,
Schmitz-Linneweber C., R. M. Maier, J. P. Alcaraz, A. Cottet, R. G. Herrmann, R. Mache,
Schmitz-Linneweber C., M. Tillich, R. G. Herrmann, R. M. Maier,
Sears B. B., L. L. Stoike, W. L. Chiu,
Shinozaki K., M. Ohme, M. Tanaka, et al. (20 co-authors)
Sriraman P., D. Silhavy, P. Maliga,
Stoebe B., K. V. Kowallik,
Stoebe B. W., Martin, K. V. Kowallik,
Stubbe W.,
Sugita M., M. Sugiura,
Sugita M., Z. Svab, P. Maliga, M. Sugiura,
Sugiura M., T. Hirose, M. Sugita,
Svab Z., P. Maliga,
Than N. D., A. Páy, M. A. Smith, P. Medgyesy, L. Márton,
Tsudzuki T., T. Wakasugi, M. Sugiura,
Vera A., M. Sugiura,
Weihe A., T. Börner,
Wolfe K. H., W. H. Li, P. M. Sharp,
Wolfe K. H., C. W. Morden, J. D. Palmer,
Young N. D., C. W. dePamphilis,